Evaluation of the PowerPlex® Fusion System for Use on the ABI PRISM® 310 Genetic Analyzer
Northeastern Illinois Regional Crime Laboratory, Vernon Hills, Illinois, USA
Publication Date: 2013
Introduction
The increasing need for expanded multiplex amplification systems for human identification will require changes to both chemistry and instrumentation used in the forensic DNA laboratory. Budgetary and space constraints may limit instrument upgrades in low- to medium-sample-throughput laboratories. The PowerPlex® Fusion System offers an expanded multiplex amplification system without instrument upgrades for laboratories still using the ABI PRISM® 310 Genetic Analyzer. The PowerPlex® Fusion System is a 24-locus multiplex amplification chemistry using a 5-color dye system for fluorescent detection of the 13 core CODIS loci and 12 core European Standard Set loci. Amelogenin and DYS391 loci also are present (1) (2) .
Currently, PowerPlex® Fusion protocols are available for the ABI PRISM® 3100 series and Applied Biosystems® 3130 and 3500 series of genetic analyzers (2) . We conducted the following evaluation to determine if the PowerPlex® Fusion System is suitable for use on the ABI PRISM® 310 Genetic Analyzer. We evaluated the PowerPlex® Fusion System for its reproducibility, precision and sensitivity. We also amplified case-type samples and a few unique pre-characterized case samples to demonstrate PowerPlex® Fusion’s efficacy with samples commonly seen in forensic DNA casework.
Materials and Methods
Sample Collection: Buccal swab samples were collected from each of nine volunteers from the Northeastern Illinois Regional Crime Laboratory (NIRCL) for use in reproducibility and concordance studies. These samples were extracted by robotic extraction using the QIAGEN EZ1® Advanced XL with the Investigator Protocol, quantitated with the Applied Biosystems Quantifiler® Duo and previously characterized with the Applied Biosystems AmpFlSTR® Identifiler® Plus PCR Amplification Kit.
Samples from nonprobative casework and unique cases also were extracted by either robotic (EZ1® instrument) or organic extraction, quantitated with Quantifiler® Duo and previously characterized with the AmpFlSTR® Identifiler® Plus or AmpFlSTR® Profiler Plus® and COfiler® kits.
Matrix samples were prepared using the PowerPlex® 5-Dye Matrix Standards, 310 (Cat.# DG4600). These samples were prepared using 1µl of matrix dye and 24µl of Hi-Di™ formamide (Applied Biosystems).
Amplification: Amplification for PowerPlex® Fusion was performed using the manufacturer’s recommendations and the following volumes and amplification protocol on the GeneAmp® PCR System 9700 thermal cycler (Applied Biosystems):
| Component | Volume Per Reaction |
| Water, Amplification Grade | to a final volume of 25.0µl |
| PowerPlex® Fusion 5X Master Mix | 5.0µl |
| PowerPlex® Fusion 5X Primer Pair Mix | 5.0µl |
| Template DNA (0.25–0.5ng) | up to 15.0µl |
| Total reaction volume | 25µl |
Amplification Protocol
96°C for 1 minute, then:
94°C for 10 seconds
59°C for 1 minute
72°C for 30 seconds
for 30 cycles, then:
60°C for 10 minutes
4°C soak
The 2800M Control DNA was diluted to 0.5ng/µl, and 1µl of diluted DNA was added to the reaction as a positive control.
Amplification Grade Water included in the PowerPlex® Fusion System was added to the reaction as a negative control.
Amplified DNA Detection: The following volumes were combined and loaded onto an ABI PRISM® 310 Genetic Analyzer for analysis:
| Component | Volume Per Reaction |
| CC5 Internal Lane Standard 500 | 1.0µl |
| Hi-Di™ formamide | 24.0µl |
Twenty-five microliters of this mix was combined with 1.0µl of amplified sample in each sample tube. One microliter of PowerPlex® Fusion Allelic Ladder was combined with 25µl of mix.
The following run conditions were used with the ABI PRISM® 310 Genetic Analyzer:
- Capillary Length: 47cm × 50µm
- Polymer: POP-4®
- Module: GS STR POP4 (1ml) G5v2.md5
- Injection time: 5 seconds (unless noted otherwise)
- Run time: 28 minutes
- Inj. kV: 15.0
- Run kV: 15.0
- Run °C: 60
Results were analyzed using GeneMapper® ID software, version 3.2 (Applied Biosystems). Panels and bins for PowerPlex® Fusion were downloaded from the Promega web site at: www.promega.com/resources/tools/genemapper-idsoftware-panels-and-bin-sets/. Analysis methods were set as recommended in the PowerPlex® Fusion System Technical Manual #TMD039 unless otherwise noted.
Results
Reproducibility and Concordance Studies
We performed the reproducibility study as a series of different components to evaluate the samples over multiple runs and injections.
PowerPlex® Fusion Allelic Ladder: We injected a total of twelve ladders, three times each, to evaluate the allele-calling capabilities of the PowerPlex® Fusion panel and bin sets over different sample preparations and multiple injections of the same sample. Additionally, this allowed us to evaluate the sizing interpolation of the CC5 Internal Lane Standard over different sample preparations and multiple injections. Average RFU values for each of the five dyes were at optimal levels for detection (Fluorescein, 1000RFU; JOE, 1500RFU; TMR-ET, 1000RFU; CXR-ET, 1500RFU; CC5, 700RFU). We were able to detect all intended alleles in the PowerPlex® Fusion Allelic Ladder with their proper designations.
Analytical Threshold: We injected each of six manipulation blank samples for 3, 5 and 10 seconds. For each dye, we recorded the averages of the three highest peak RFU values along with the three lowest trough values. We then calculated the difference between the peaks and troughs and multiplied these values by two (SWGDAM Interpretation Guidelines, 2010). These values are shown in Table 1.
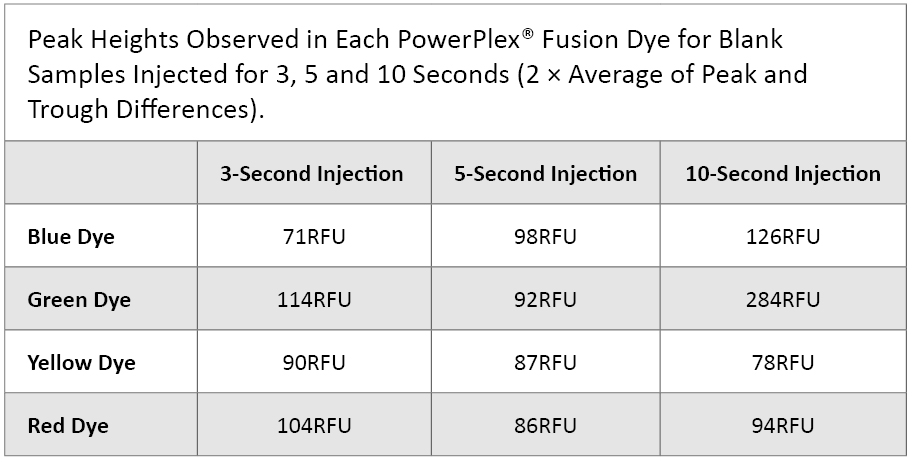 Table 1. Fluorescence Levels Observed in Each PowerPlex® Fusion Dye for Blank Samples Injected for 3, 5 and 10 seconds.
Table 1. Fluorescence Levels Observed in Each PowerPlex® Fusion Dye for Blank Samples Injected for 3, 5 and 10 seconds. Most of the peak heights fell below 100RFU. However, different analytical thresholds could be set for each dye or different injection times if further data support this trend.
Concordance: We collected buccal swab samples from each of nine volunteers from the NIRCL. These samples were previously characterized with the Identifiler® Plus kit. All of the alleles amplified with the PowerPlex® Fusion System for each sample correctly corresponded to allele calls in the respective loci with the Identifiler® Plus system. Additionally, we observed an allele at the DYS391 locus for all male profiles; no alleles were observed for female profiles.
We observed some reproducible artifacts at approximately 214.55bp and 247.97bp in the green dye and 172.54bp in the orange dye. Many of these fell below the 100RFU analytical threshold. However, in a few instances, they were above the 100RFU analytical threshold. These correspond to artifacts documented by Promega in the PowerPlex® Fusion System Technical Manual.
Precision Study
The PowerPlex® Fusion Allelic Ladder samples previously analyzed for the reproducibility study were evaluated for precision. We tabulated base pair sizes for eleven different alleles and one haplotype. These represent three different loci in each of the dyes. Averages and standard deviations are shown in Table 2.
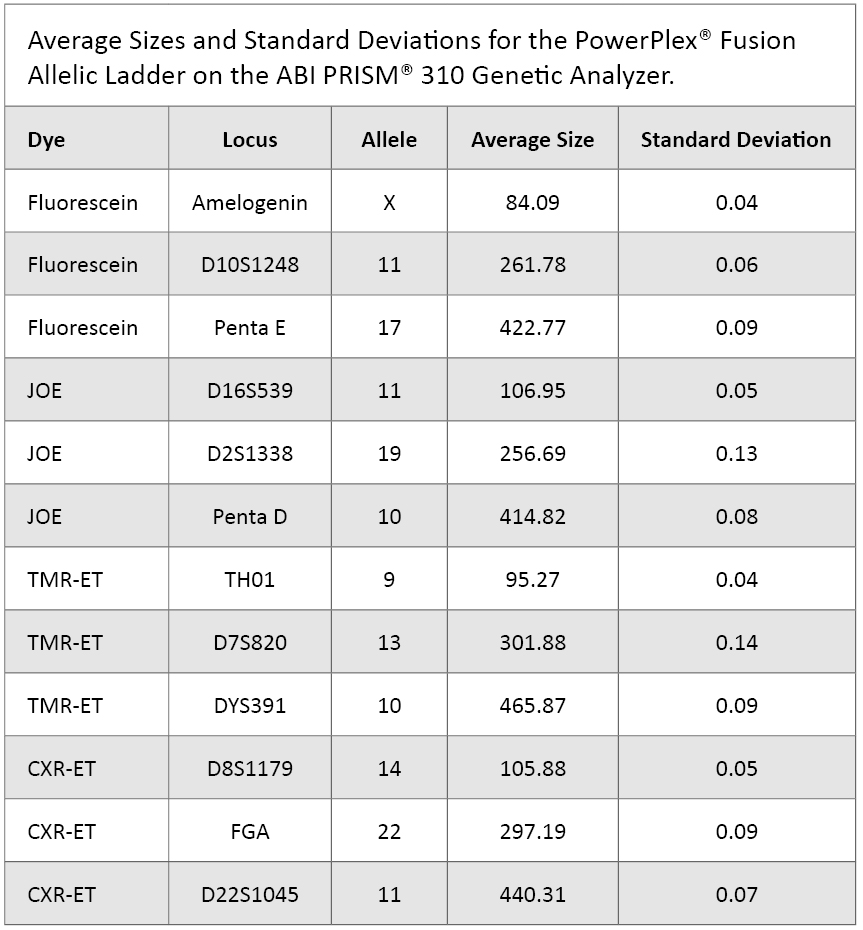 Table 2. Average Sizes and Standard Deviations for the PowerPlex® Fusion Allelic Ladder on the ABI PRISM® 310 Genetic Analyzer.
Table 2. Average Sizes and Standard Deviations for the PowerPlex® Fusion Allelic Ladder on the ABI PRISM® 310 Genetic Analyzer. Sensitivity Study
We diluted the 2800M Control DNA stock solution of 10ng/µl to the recommended concentration of 500pg in the appropriate template volume for amplification. We then amplified serial dilutions of the 500pg sample using PowerPlex® Fusion to evaluate levels of detection and allelic dropout. Table 3 reflects these results.
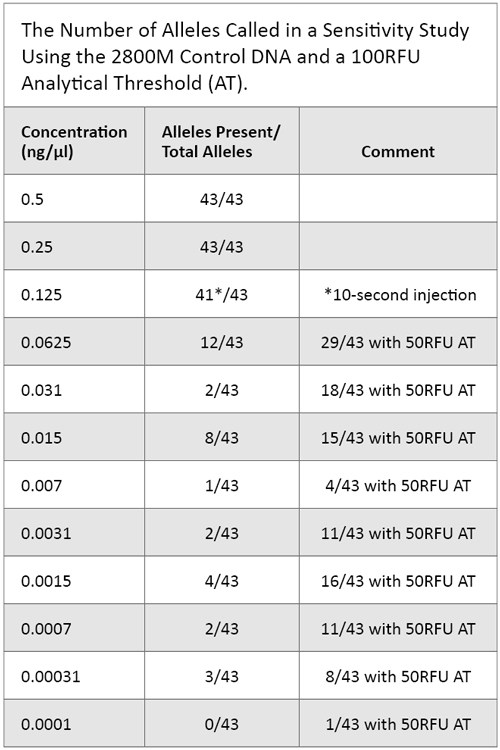 Table 3. The Number of Alleles Called in a Sensitivity Study Using the 2800M Control DNA and a 100RFU Analytical Threshold (AT).
Table 3. The Number of Alleles Called in a Sensitivity Study Using the 2800M Control DNA and a 100RFU Analytical Threshold (AT). Nonprobative Casework and Unique Cases
We quantitated 28 different nonprobative casework samples previously extracted by robotic method or organic method and previously characterized by AmpFlSTR® Profiler Plus®/COfiler® or Identifiler® Plus kits and Quantifiler® Duo, then amplified the DNA with the PowerPlex® Fusion System. We obtained consistent results for the 28 samples, with one exception at the D21S11 locus (Figure 1). For this sample, the initial Profiler Plus® result was called an off-ladder allele (OLA) and a 30. Upon comparison to the Profiler Plus® allelic ladder, a 29.3 was calculated for the OLA. The PowerPlex® Fusion result indicated a split peak; however, only a 30 allele was called.
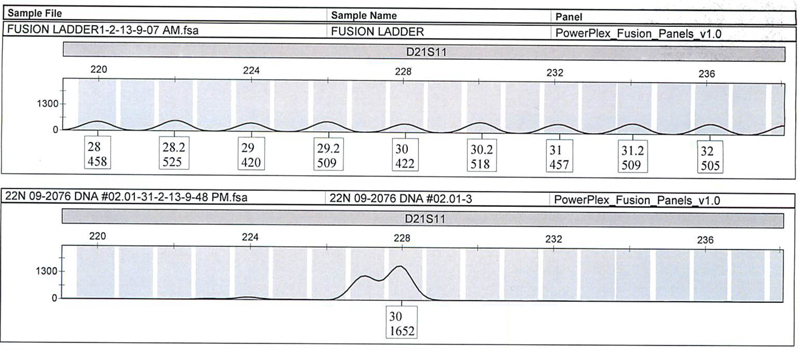 Figure 1. Expanded view of D21S11 locus of the PowerPlex® Fusion Allelic Ladder (top panel) and buccal swabs standard (bottom panel) with 5-second injections.
Figure 1. Expanded view of D21S11 locus of the PowerPlex® Fusion Allelic Ladder (top panel) and buccal swabs standard (bottom panel) with 5-second injections. We also amplified some unique samples discovered in casework with the PowerPlex® Fusion System. These are as follows:
Unique Sample 1: We obtained a tri-allelic profile in an Identifiler® Plus amplification at the D21S11 locus. When the same sample was amplified with PowerPlex® Fusion, we observed the same tri-allelic profile.
Unique Sample 2: An OLA was detected in an Identifiler® Plus amplification. The OLA fell at the far left edge of the D13S317 locus bins. The Identifiler® Plus Allelic Ladder at D13S317 includes alleles 8–15; therefore the allele was called <8 according to NIRCL protocols. When the sample was amplified with the PowerPlex® Fusion System, a 5 allele was called.
Discussion
After an initial evaluation, our results indicate that the PowerPlex® Fusion System is a very sensitive and robust system and would be suitable for use on the ABI PRISM® 310 Genetic Analyzer. A laboratory with limited budget and lower throughput would find this combination of kit and instrumentation an alternative to a larger instrument upgrade.
The reproducibility study indicated that the run parameters and reagent formulations were suitable for accurate results. We found that the analytical threshold was instrument- and dye-dependent; however, the analytical threshold established was not significantly different from the one currently in use.
The precision study demonstrated how little variation there is in the alleles from run to run. We expected a multiplex system of this size to have some variation, especially in the loci containing the larger amplicons. This was not observed. We specifically chose loci in each dye by different fragment sizes to determine if any variation existed. We observed the greatest standard deviations in D2S1338 and D7S820 (0.13 and 0.14, respectively). These loci are located within the middle of the size range and were not far off the standard deviations of the other loci evaluated.
The sensitivity study indicated that 500–250pg is the amount of DNA needed to obtain a complete profile. This range corresponds to the amount recommended in the PowerPlex® Fusion System Technical Manual. We noted in the present study that 125pg produced a nearly complete profile with a 10-second injection; however, as observed in the analytical threshold study, increased injection time may increase artifacts. Sensitivity samples below 125pg showed significant allelic dropout; however, we still obtained useful information with a lower analytical threshold. We did not perform mixture and stochastic studies in the current study, but the lower DNA concentrations did indicate preferential and differential amplification.
The final study examined nonprobative samples previously characterized in other amplifications to determine concordance and the ability of the PowerPlex® Fusion System to amplify samples routinely seen in casework. The samples were concordant between PowerPlex® Fusion and AmpFlSTR® Profiler Plus®/COfiler® and AmpFlSTR® Identifiler® Plus.
Acknowledgments
The Northeastern Illinois Regional Crime Laboratory would like to thank Promega Corporation for the invitation to evaluate the PowerPlex® Fusion System on the ABI PRISM® 310 Genetic Analyzer.
Article References
- Oostdik, K. et al. (2012) Bridging databases for today and tomorrow: The PowerPlex® Fusion System. Profiles in DNA.
- PowerPlex® Fusion System Technical Manual , TMD039, Promega Corporation..
How to Cite This Article
Scientific Style and Format, 7th edition, 2006
Pfoser, K. and Owen, S. Evaluation of the PowerPlex® Fusion System for Use on the ABI PRISM® 310 Genetic Analyzer. [Internet] 2013. [cited: year, month, date]. Available from: https://www.promega.com/resources/profiles-in-dna/2013/evaluation-of-the-powerplex-fusion-system-for-use-on-the-abi-prism-310-genetic-analyzer/
American Medical Association, Manual of Style, 10th edition, 2007
Pfoser, K. and Owen, S. Evaluation of the PowerPlex® Fusion System for Use on the ABI PRISM® 310 Genetic Analyzer. Promega Corporation Web site. https://www.promega.com/resources/profiles-in-dna/2013/evaluation-of-the-powerplex-fusion-system-for-use-on-the-abi-prism-310-genetic-analyzer/ Updated 2013. Accessed Month Day, Year.
Contribution of an article to Profiles in DNA does not constitute an endorsement of Promega products.
PowerPlex is a registered trademark of Promega Corporation.
ABI PRISM, Applied Biosystems, COfiler, GeneAmp, GeneMapper, Identifiler, Profiler Plus, Quantifiler are registered trademarks of Applied Biosystems. AmpFlSTR is a registered trademark of Applera Corporation. EZ1 is a registered trademark of Qiagen GmbH Corporation. Hi-Di is a trademark of Applera Corporation. POP-4 is a registered trademark of Life Technologies Corporation.